<< back to search
IMAGES:
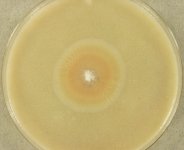
Search Details
add to cart
| UAMH Number: | 10459 |
|---|---|
| Species Name: | Oidiodendron fimicola |
| Type: | Oidiodendron fimicolum |
| Synonyms: | |
| Taxonomy: | FUNGI Ascomycota, Leotiomycetes, Helotiales, Myxotrichaceae |
| Strain History: | D. Bayer (DC 60) -> Currah, R.S. -> UAMH |
| Substrate: | mushroom compost | Location: | USA Missouri, St. Louis (GEO: 38.627,-90.199) |
| Isolator: | D. Bayer (DC 60) |
| Isolation Date: | 1976 |
| Date Received: | 2004-08-17 |
| Characters: | APPLICATION use of Biolog profiles to distinguish Oidiodendron species - Rice A, Currah RS, Stud Mycol 53:75-82, 2005 // MOLECULAR SYSTEMATICS Blasts among Oidiodendron species // MOLECULAR SYSTEMATICS ITS sequence - fide UAMH 2011 |
| Compounds: | |
| Cross Reference: | |
| Pathogenic Potential: | Human: no | Animal: no | Plant: yes |
| Biosafety Risk Group: | RG2 (check the PHAC ePATHogen Risk Group Database for updates) |
| Regulatory Requirements: | Canadian requesters must provide PHAC Pathogen and Toxin License Number (see: https://www.canada.ca/en/public-health/services/laboratory-biosafety-biosecurity/licensing-program.html) and a CFIA Written Authorization for transfer (http://www.inspection.gc.ca/plants/plant-pests-invasive-species/directives/date/d-12-03/eng/1432656209220/1432751554580#app2) prior to shipment. International requesters must provide all legally required importation documentation prior to shipment. This strain is not available for shipment to Cuba, the Democratic People's Republic of Korea, Iran or Syria. Plant pathogenicity status may be verified by using the USDA Agricultural Research Service (ARS) Fungal Database |
| MycoBank ID: | 500255 |
| Sequences: | >UAMH10459_ITS TCATTACAGAGTTCTCGCCCTCCTGGGTAGATCTCCCATCTACTGTGATTGCTACCGCGTTGCTTTGGCGGGCCGCCGGGCCCTGCCCGCCGCCGGCCCCGGCCGGCGCGCGCCCGCCAGAGGCCCCTAAACCCGAATGTCTAGTGTCGTCTGAGCAGCACATAATCGTTAAAACTTTCAACAACGGATCTCTTGGTTCTGGCATCGATGAAGAACGCAGCGAAATGCGATAAGTAATGCGAATTGCAGAATTCAGTGAGTCATCGAATCTTTGAACGCACATTGCGCCCCGTGGTATTCTGCGGGGCATGCCTGTTCGAGCGTCATTTCAACCCTCAAGCTCCGCTTGGTGTTGGGCCGGGCCCGTCGCGGCCGGCCCCAAAGACAGTGGCGGCGCCGCCTGGCTCTCAGCGTAGTACCTCT |
IMAGES:
